Do you like long reads and BAM?
| 5 May, 2017 | Hollydawn Murray |
|

|
For the final day of London Calling 2017, Hollydawn Murray, shares the focus for this year's conference and summarises the technical and software updates for Nanopore technology
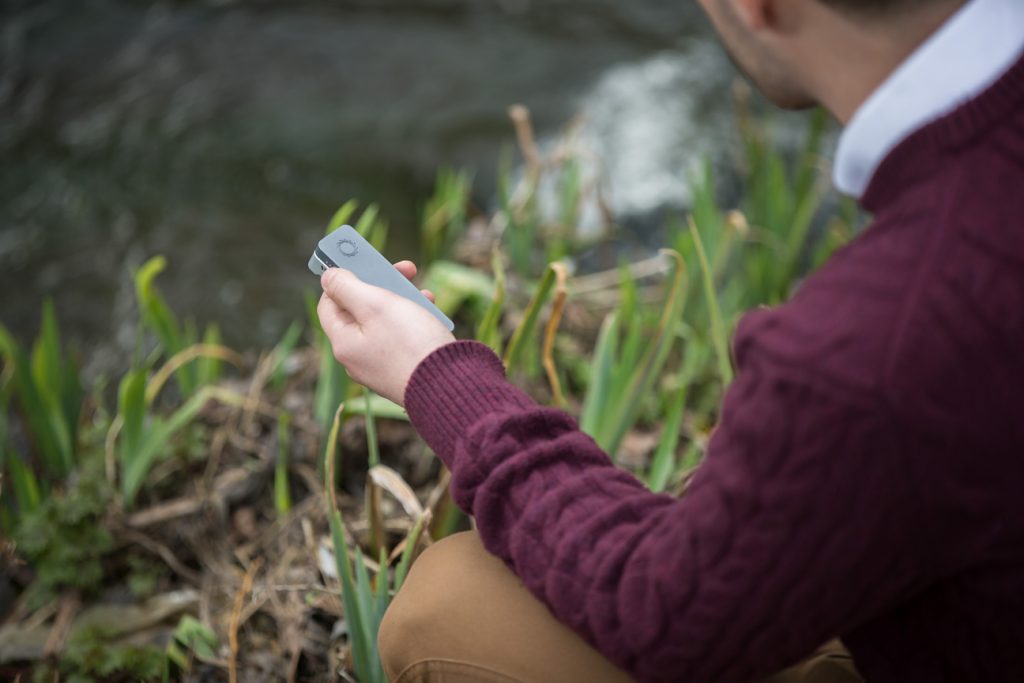

Credit: Oxford Nanopore Technologies
Today marks the second and final day of London Calling 2017, a conference dedicated to the latest Nanopore genetic-sequencing technology, such as the portable MinION, and is known for its excitement.
The megabase read remains elusive. But that is not to say that Nanopore technology does not continue to evolve quickly, after all more than just size matters.
As predicted by Keith Robison earlier this week on his Omics! Omics! Blog, there has been a particular focus on long reads – reads approaching 1 megabase – with the term featuring prominently in the titles of talks by Josh Quick (University of Birmingham) and Kazuharu Arakawa (Keio University), workshops led by Miten Jain (University of California, Santa Cruz) and Hans Jansen (ZF-screens B.V.), and a plenary delivered by Karen Miga (University of California, Santa Cruz). Still, the megabase read remains elusive. But that is not to say that Nanopore technology does not continue to evolve quickly, after all more than just size matters.
This week, in an article published in our Nanopore channel, Hans Jansen (ZF-screens B.V.) and colleagues reported improvements in the R9 chemistry – an alternative nanopore chemistry released in a new flow cell – that are essential to the assembly of the mitochondrial genome, in comparison to the R7.3 chemistry.
David Eccles (Malaghan Institute of Medical Research) and co-authors also used the improved R9 flow cells (R9.4 to be exact) in their paper titled ‘Investigation of chimeric reads using the MinION’ published today on F1000Research; they found that chimeric reads may be present in the output of sequencing runs. Given the minute half-life of developments in this field, it is not surprising that their discussion ends with a tip off to the imminent release of R9.5 chemistry and 1D2 base calling.
At this point, technical and software updates have come to be the norm as Oxford Nanopore Technologies pushes towards its goal “to enable the analysis of any living thing, by any person, in any environment”. And given that the phase 1 data release was only published 18 months ago, the community appears to be steadily progressing to this end. Here’s a kb-scale summary:
The portability of the MinION means sequencing is no longer tethered to the lab
Would you like them with a mouse?
From roundworms to red algae, Nanopore has been useful in sequencing the DNA of a variety of species. But Nanopore sequencing is not just limited to DNA, nor is it only about whole genome sequencing. Mark Akeson’s lab (University of California, Santa Cruz) recently sequenced bacterial ribosomal RNA (no, not the cDNA) directly and the reaction from the community did not mince words: Nick Loman said the work was “too awesome ”. This is especially timely given that, as of only a few weeks ago, direct RNA sequencing kits are now available to all MinION users.
Would you, could you, on a boat?
The portability of the MinION means sequencing is no longer tethered to the lab, and it seems some researchers have taken this as a challenge! Not only has the MinION seen snow and (presumably) rain, it has also been used to successfully sequence DNA in-flight on the International Space Station. Recently, Edwards (Aberystwyth University) et al. went in the opposite direction. In their preprint, the group literally took Nanopore to new depths showing that the MinION works 100m underground in a coal mine, which demonstrated its potential for on-site metagenomics. For a first-hand look, watch this video.
In a car?
Open sharing of the genetic data has helped to build a better understanding of Zika phylogeny, transmission and diversity
And while space is impressive, admittedly, my favourite Nanopore application involves a road trip in Brazil. Almost certainly less glamorous than it sounds, the ZiBRA (Zika in Brazil in Real Time Analysis) project sequenced Zika virus genomes from patients with a range of clinical presentations, and from a broad geographical region by way of a mobile laboratory.
Open sharing of the genetic data has helped to build a better understanding of Zika phylogeny, transmission and diversity, as visualised by NextStrain. Similarly, field based surveillance has been previously employed during an Ebola outbreak.
And while it may go without saying, there is also an expanding list of clinical applications (such as the same-day diagnosis of tuberculosis) and the potential for antimicrobial resistance testing (for instance, in bacterial isolates and in urine). Considering the exceptional progress made thus far, it’s difficult to imagine what excitement London Calling 2018 holds!
I do like long reads and BAM.

|


This piece of writing provides clear idea in favor of the new people of blogging, that genuinely how to do
blogging.